FixedLengthSpiral
- class picazzo3.wg.spirals.cell.FixedLengthSpiral
Spiral with incoupling sections that calculates its length. The total length is set by the property
total_lengthand the inner size of the spiral will be adapted so that the total length of the spiral (including the incoupling sections) would be equal tototal_length. The way this inner size is calculated can set using properties in the Layout view.- Parameters:
- total_length: float and number > 0
Total design length of the spiral.
- n_o_loops: int and number > 0
Number of loops in the spiral
- trace_template: PCell and _TraceTemplate
Trace template used in the chain.
- name: String that contains only ISO/IEC 8859-1 (extended ASCII py3) or pure ASCII (py2) characters
The unique name of the pcell
- Other Parameters:
- n_o_traces: int and number > 0, locked
Total number of traces used in the spiral.
- traces: List with type restriction, allowed types: <class ‘ipkiss3.pcell.cell.pcell.PCell’>, locked
Views
- class Layout
The inner size of the spiral is calculated by assuming a minimal inner_size and growing it in either the direction set by
growth_direction. An error is raised when that is impossible to do with the set number of loops.- Parameters:
- growth_direction: List with value restriction, allowed values: [‘H’, ‘V’]
- incoupling_length: float and Real, number and number >= 0
length of the incoupling section.
- shapes: list
List of shapes used to build the traces
- flatten: ( bool, bool_ or int )
If true the instances are flattened
- view_name: String that contains only alphanumeric characters from the ASCII set or contains _$. ASCII set is extended on PY3.
The name of the view
- stub_direction: List with value restriction, allowed values: [‘H’, ‘V’]
- spacing: float and Real, number and number >= 0
spacing between the individual loops.
- Other Parameters:
- auto_transform: locked
- inner_size: locked
- spiral_center: locked
Examples
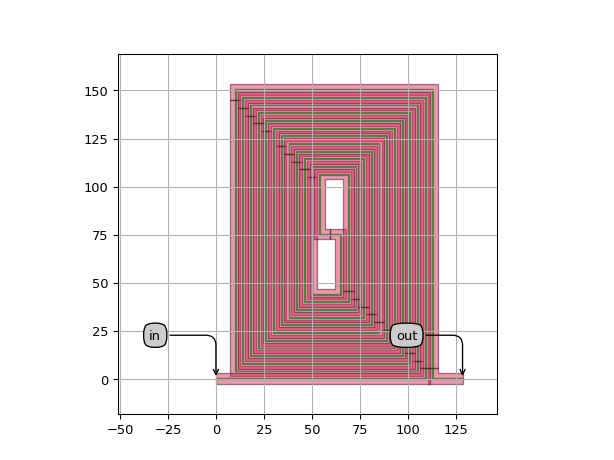
import si_fab.all as pdk # noqa: F401 from picazzo3.wg.spirals import FixedLengthSpiral from ipkiss3 import all as i3 cell = FixedLengthSpiral(total_length=4000, n_o_loops=6, trace_template=i3.TECH.PCELLS.WG.DEFAULT) layout = cell.Layout( incoupling_length=10.0, spacing=4, stub_direction="H", # either H or V growth_direction="V", # either H or V ) # Checking if the trace length is indeed correct print(layout.trace_length()) layout.visualize(annotate=True)